Plagiarized Errors and Molecular Genetics
Another argument in the evolution-creation controversy
by Edward E. Max, M.D., Ph.D.The following essay is an updating of an article I published in Creation/Evolution in 1986 (XIX, p.34). I am posting it with permission from Creation/Evolution.
![]()
|
Other Links:
|
1. Introduction
Most scientists regard the evidence for evolution as overwhelming. Thus, in their conviction that evolution has already been thoroughly and sufficiently documented, they sometimes fail to consider how new discoveries might provide evidence for evolution that might be powerfully persuasive to individuals leaning towards creationist beliefs. In this article, I describe some discoveries from my own field of molecular genetics, discoveries whose implication for the creation-evolution controversy were not explicitly discussed when they were reported. I try to show how they provide evidence for evolution that is both convincing and conceptually simple enough for the interested layperson to appreciate.
The new molecular evidence bears on a question which, in my opinion, represents one of the few cases in which a creationist argument had demonstrated logical consistency and had fought the evolutionary position to a deadlock. This is the question of how to interpret the similarities between modern living species, especially the similarities observed at the molecular level. As we will see, the recent discoveries from molecular genetics resolve this deadlock unequivocally in favor of evolution.
1.1 The evolutionary view of species similarities
Consider first how evolutionists interpret similarities between species living today. Present-day humans and chimpanzees, despite obvious external and behavioral differences, have extremely similar internal organs and physiological functions; indeed their genes are more than 98% identical (Goodman et al., J Molec Evolution 30:260,1990). Just as the resemblance between two siblings suggests a common parentage, resemblance between species suggests common ancestors. Evolutionists believe that humans, gorillas, and chimpanzees evolved from a common ancestor: an ape-like creature that lived perhaps five to ten million years ago, rather recently on the geological time scale. (The thought that humans and apes might share a common ancestor seems particularly unacceptable to creationists because of the theological implications of such a relationship and the clear contradiction to the creationists' literal interpretation of biblical Genesis.) Species less similar to humans than are apes--mice, for example--are believed to have branched off millions of years earlier from a common primitive mammalian ancestor. Evolutionary family tree diagrams that express such relationships between species have been constructed by evolutionary biologists by analyzing similarities of present-day organisms. In many cases, fossilized remains of extinct species can be used to support the features of such evolutionary trees; fossil evidence will not, however, be discussed in this article.
Another extensive source of data that has been of major importance in constructing similarity tree diagrams is the species comparison of proteins and genes. Proteins are large biological molecules made of subunits called amino acids that are attached to one another in chains, like the cars of a train. There are twenty different kinds of amino acids used in proteins, and most proteins contain hundreds of these subunits. Each protein has a specific number and sequence of amino acids, and this sequence determines what properties that protein will have. In order for a cell to synthesize a specific protein, it must access an "information bank" in which amino acid sequences are stored; this information bank is comprised of the organism's genes, which contain the amino acid sequences encoded in molecules of deoxyribonucleic acid (DNA). Biochemists can purify proteins and learn the exact sequence of their amino acids, or they can obtain this information by reading the appropriate sequence from an organism's DNA. Considerable effort has gone into comparing the sequences of similar proteins isolated from different species. For example, one protein called "cytochrome c" has been examined in more than eighty species. These cytochrome c amino acid sequences represent "digital" bits of data that can be used to quantify differences between species, and these differences can be used to construct evolutionary trees much like those based on comparisons of "analog" features of body anatomy. Such protein sequence trees--as well as trees based upon DNA structure similarities--agree remarkably well with the evolutionary trees derived earlier from anatomic similarities. The agreement of evolutionary trees constructed from such completely different sorts of data (e.g. Goodman et al Mol Phylogenet Evol 9:585, 1998) has been taken by evolutionists as evidence of the validity of the intellectual framework on which the trees are based: the theory of evolution (see Jukes, in Scientists Confront Creationism, edited by Godfrey, WW Norton, New York 1983; Creation/Evolution XVIII:42, 1986; Goodman et al., J Mol Evol 30: 260, 1990).
1.2 The creationist view of species similarities leads to a deadlock
However, creationists have an alternative interpretation of the amino acid sequence similarities reflected in the evolutionists' trees. They say that such sequence similarities in "related" species simply reflect the creator's choice to design similar species to function similarly, not only at the level of bones, muscles and organs, but also at the level of protein function--hence the amino acid sequence similarities.
Thus the similarities between species in anatomy and protein structure can be interpreted in two entirely different ways. The evolutionists say that the similarity between features of, for example, humans and apes reflects the fact that these features were inherited from a common ancestor; that is, the similar features of humans and apes are determined by modern copies of genes that once existed in species that was ancestral to both apes and humans. The creationists say that apes and humans were created independently but were designed with similar features so that they would function similarly. Both the gene copying and the independent creation views seem consistent with the similarity data, but which view is correct?
1.3 A possible rationale to resolve the deadlock
One way to distinguish between copying and independent creation is suggested by analogy to the following two cases from the legal literature. In 1941 the author of a chemistry textbook brought suit charging that portions of his textbook had been plagiarized by the author of a competing textbook (Colonial Book Co, Inc. v. Amsco School Publications, Inc., 41 F. Supp.156 (S.D.N.Y. 1941), aff'd 142 F.2d 362 (2nd Cir. 1944)). In 1946 the publisher of a trade directory for the construction industry made similar charges against a competing directory publisher (Sub-Contractors Register, Inc. v McGovern's Contractors & Builders Manual, Inc. 69 F.Supp. 507, 509 (S.D.N.Y. 1946)). In both cases, mere similarity between the contents of the alleged copies and the originals was not considered compelling evidence of copying. After all, both chemistry textbooks were describing the same body of chemical knowledge (the books were designed to "function similarly") and both directories listed members of the same industry, so substantial resemblance would be expected even if no copying had occurred. However, in both cases errors present in the "originals" appeared in the alleged copies. The courts judged that it was inconceivable that the same errors could have been made independently by each plaintiff and defendant, and ruled in both cases that copying had occurred. The principle that duplicated errors imply copying is now well established in copyright law. (In recognition of this fact, directory publishers routinely include false entries in their directories to trap potential plagiarizers.)
Can "errors" in modern species be used as evidence of "copying" from ancient ancestors? In fact, the answer to this question appears to be "yes," since recent molecular genetics investigations have uncovered some examples of the same "errors" present in the genetic material of humans and apes. To understand these findings it is necessary to know a little about DNA, the chemical molecule in which genetic information is stored.
2.1 DNA Basics
In one respect the basic structure of DNA resembles that of proteins: both are made of linear chains of varying subunits. Apart from this common feature, DNA structure is quite different from that of proteins. The subunits in DNA are called nucleotides or bases, and the sequence of these nucleotides contains the genetic information specifying the sequence of amino acids in each protein made by the organism. Whereas 20 different amino acids comprise the subunits of proteins, there are only four different nucleotide bases in DNA, generally abbreviated A, T, G and C. According to the "genetic code" deduced by scientists in the 1960s, each amino acid is specified by one or more triplets of nucleotides; for example, the sequence GCG specifies the amino acid alanine. Since there are 64 different triplets (each called a codon) and only 20 amino acids to specify, some amino acids are represented by more than one triplet (e.g. ATA, ATC and ATT all code for the amino acid isoleucine); and three triplets -- TAA, TAG and TGA -- represent "stop codons" that mark the end of the gene sequence that can be used to specify amino acid sequence.
|
|
|
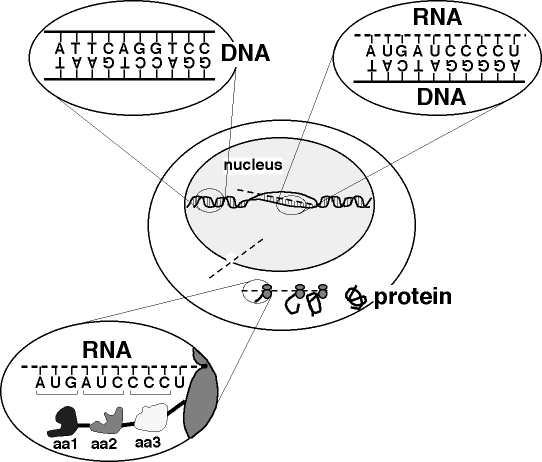
Figure 1. DNA Basics. The central oval represents a cell, within which lies the nucleus. Inside the nucleus, most of the DNA exists as a double helix. The oval at upper left shows an expanded view of the DNA, in which the helices have been drawn "untwisted" to reveal similarity to a ladder. The genetic information is stored in the sequences of nucleotide bases (A, T, G or C) that form the rungs of the ladder. Each rung is formed by a pair of nucleotide bases touching each other, one base attached to one strand backbone, and the other attached to the other strand backbone. An "A" nucleotide always pairs with a "T," and a "G" always pairs with a "C." In order to synthesize a protein, the cell reads the genetic information of the gene for that protein by "transcribing" a molecule of RNA from the gene. For transcription, the strands of the DNA double helix must partially separate so that the bases that form RNA can assemble according to the rules of complementary basepairing. The expanded view at upper right shows the two major differences between RNA and DNA: the RNA backbone strand has a slightly different chemical structure (represented by the dashed line), and a slightly modified form of "T" known as "U" is found in RNA. The transcribed strand of RNA acts as a "messenger" that carries the genetic information from storage in the nucleus to the protein manufacturing modules (represented in the figure by double grey ovals) in the cytoplasm. The expanded view at lower left shows that the sequence of RNA bases is read so that each triplet of bases specifies an amino acid (aa1, aa2, etc.) in the protein. The protein folds into a functional three-dimensional structure that depends on the linear sequence of amino acids. |
DNA contains two linear chains in a double-stranded structure that resembles a twisted ladder--the famous "double helix." The vertical beams of the ladder represent a uniform backbone chain which contains no sequence information. As shown in Figure 1 above, the information is stored in the "rungs" of the ladder, which are formed from a pair of nucleotide bases, each sticking out from one vertical backbone strand and touching the base from the opposite strand to form a "rung." The base G on one strand always contacts a base C on the opposite strand; similarly an A always contacts a T. Thus a string of Ts on one strand can "basepair" or "anneal" with a strand containing a string of As to form a double-stranded structure. The sequence of nucleotide bases in one strand is said to be "complementary" to the sequence of the other strand. For any one gene the triplets of bases encoding amino acid sequence are on only one strand. Some genes are encoded on one strand, while other genes lie on the other strand. In most mammalian genes the DNA coding for amino acid sequences is interrupted by segments of apparently meaningless DNA ("introns"). Intron sequences need to be removed before the sequence is used to assemble amino acids; this removal, or splicing, does not occur in the DNA molecule, but in the next stage of information transfer.
In order for a cell to produce a particular protein whose amino acid sequence is encoded in a gene, the sequence information in the DNA must first be copied or "transcribed" into a single-stranded molecule called ribonucleic acid (RNA), as shown above in Figure 1. This initial transcript of RNA undergoes several structural alterations, known collectively as "processing," before it is used to assemble amino acids. These processing steps include the "splicing" out of unnecessary intron segments from the RNA and the addition of nucleotides at one end--the "poly(A) tail"--which promote proper functioning of the RNA in the cell. It is the "processed" RNA that participates directly in the assembly of amino acids into proteins. The transcription of a gene into an RNA copy is very tightly controlled, in part by highly specific regulatory sequences known as promoters that for most genes occur in the DNA just outside the transcribed region but close to the position where the transcription into RNA should start.
When a cell divides, the entire sequence of its DNA must be duplicated into two faithful copies of the original; one copy goes to each of the "daughter" cells created by the division. Occasionally, errors occur in this copying mechanism, creating "mutations" in the DNA sequence. There are several types of mutations, including substitutions of one or a few nucleotides, deletions of nucleotides, duplication of segments of DNA or insertion of extraneous DNA segments into an unrelated DNA sequence. Such changes can occur in most cells in the body--liver, skin, muscle, etc.--without being transmitted to offspring when the organism reproduces. However, when mutations occur in the egg or sperm or, more generally in "germline cells" (i.e., the egg or sperm plus their embryological precursors), they can be passed on to future generations. Often, mutations are inconsequential: e.g. they may fall outside a gene, or if within a gene they may not change the amino acid encoded. Many genetic differences between closely related species are thought to represent such random inconsequential mutations. Sometimes, however, mutations critically damage the function of a gene. Indeed, such mutations are the cause of genetic diseases like cystic fibrosis, sickle cell anemia, phenylketonuria, and hundreds of others, as well as many genetic aberrations studied in laboratory animals. When molecular geneticists examine the DNA of patients with such well-characterized diseases, they can almost always find the defective gene and identify the mutation that inactivated it, since it is rare for such genetic disease to be caused by a deletion that removes an entire gene. Mutations causing genetic diseases and malformations are generally so detrimental to the organism's survival and reproductive success that in the wild--i.e. in the absence of modern medical science--they would tend to be "weeded out" by the pressure of natural selection. Rarely, mutations can be beneficial to an organism: these rare cases form the basis for evolutionary adaptations that improve the "fitness" of an organism to its environment.
2.2 DNA errors
Recombinant DNA technology has in recent years allowed scientists to determine the sequence of nucleotides in segments of DNA from many species, and several billion nucleotides' worth of information has accumulated. These sequences have vastly increased our understanding of how genes normally function; but, more to the point of this article, they have provided a treasure trove of genetic "errors" that are potential clues to the analysis of copying discussed earlier. In this context I use the word "error" to include any DNA feature that we have good reason to believe (1) originated from a genetic "accident"; (2) serves no benefit to the organism carrying the features; and (3) therefore cannot reasonably be interpreted as having been "designed." I will discuss several overlapping classes of these "errors," which argue for evolution in slightly different ways. One class includes "pseudogenes," or damaged non-functional copies of genes. I will discuss three classes of pseudogenes, the last of which overlaps with another larger category of genetic "errors" known as retroposons, which will also be discussed. See Figure 2.
|
|
|
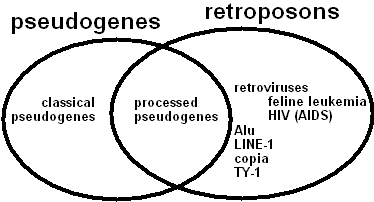
Figure 2. This Venn diagram illustrates the classes of "errors" discussed in this posting, except that the class of "unitary pseudogenes" (which is really a tiny subset of "classical pseudogenes") is not shown in the diagram. Processed pseudogenes represent the intersection of the set of pseudogenes and the set of retroposons. |
2.2.1 Pseudogenes
a. Unitary pseudogenes.
Guinea pigs and primates, including humans, get sick unless they consume ascorbic acid in their diet. For humans and guinea pigs, ascorbic acid is thus a vitamin (vitamin C), while most other species can synthesize their own ascorbic acid and thus do not require this molecule in their diet. The reason humans and guinea pigs cannot manufacture their own ascorbic acid is that they lack a functional gene encoding the enzyme protein known as L-gulono-gamma-lactone oxidase (GLO), which is required for synthesizing ascorbic acid. In most mammals functional GLO genes are present, inherited - according to the evolutionary hypothesis - from a functional GLO gene in a common ancestor of mammals. According to this view, GLO gene copies in the human and guinea pig lineages were inactivated by mutations. Presumably this occurred separately in guinea pig and primate ancestors whose natural diets were so rich in ascorbic acid that the absence of GLO enzyme activity was not a disadvantage--it did not cause selective pressure against the defective gene.
Molecular geneticists who examine DNA sequences from an evolutionary perspective know that large gene deletions are rare, so scientists expected that non-functional mutant GLO gene copies--known as "pseudogenes"--might still be present in primates and guinea pigs as relics of the functional ancestral gene. In contrast, Creationists believe that humans and guinea pigs were each created independently of all other species and must have been "designed" to function without GLO. If this were true, these two species would not be expected to carry a defective copy of the GLO gene. In fact, GLO pseudogenes have been detected in both guinea pigs and humans (Nishikimi et al. J Biol Chem 267: 21967, 1992; Nishikimi et al. J Biol Chem 269:13685, 1994), consistent with the evolutionary view; presumably, related pseudogenes also exist in non-human primates that require dietary vitamin C. The kinds of mutations found in the human and guinea pig pseudogenes are typical of the ones seen in genetic diseases like those mentioned earlier. In this essay I call the human and guinea pig GLO DNA sequences "unitary pseudogenes" to distinguish them from two other kinds of pseudogene occurring in a species that also possesses a functional copy of the same gene (see below). Readers should note that the term "unitary pseudogene" is used here for convenience; there is no standard nomenclature to describe this rare type of pseudogene.
Unitary pseudogenes are relatively rare; each is like a genetic defect that affects all individuals in a species. But these defective genes do not correspond to genetic diseases because if they caused significant symptoms or other disadvantage to their owners, individuals with intact genes would have long ago won the competition for survival and reproduction, thus driving the pseudogene out of existence. In an evolutionary perspective unitary pseudogenes represent genetic relics of genes whose function was important in ancestral species but became unnecessary in the modern species; i.e. they are vestigial DNA sequences. The presence of non-functional pseudogene relics is easily explained by the evolutionary model: they are a natural consequence of mutations that fail to be eliminated by natural selection because the function of the gene product has become unnecessary. In fact, this model predicts that many unitary pseudogenes should be found if scientists examine the genes specifying vestigial structures--for example, genes encoding eye structures in blind species such as moles or cave-dwellers. (An example confirming this prediction was recently described in marsupial moles: an apparent unitary pseudogene related to the interphotoreceptor retinoid binding protein gene [Springer et al., PNAS 94:13754, 1997]. A conceptually similar example in human DNA is provided by odorant receptor DNA sequences: about 70% are of these sequences are pseudogenes, reflecting the nearly vestigial status of our olfactory perception in comparison to that of other species [Rouquier et al., Nat Genet 18:243,1998; Rouquier, et al. Human Molec Genet &:1337,1998;Sharon et al., Genomics 61:24,1999; Rouquier et al., PNAS 97:2871,2000; Glusman et al, Genome Res 11:685, 2001].) In contrast, such pseudogenes would not be expected if each species were independently created by an intelligent designer (unless that designer were intentionally simulating evolution). Several other unitary pseudogenes are known in humans, including sequences homologous to [i.e, similar to and thought to be derived from a common ancestor] the genes encoding urate oxidase (Yeldandi et al, Gene 109:2821, 1994; Wu et al, J Mol Evol 34:78, 1992), alpha-1,3-galactosyltransferase (Galili and Swanson, PNAS 88:7401,1991) and the RT6 surface protein, a glycophosphatidylinositol-linked ADP-ribosyltransferase (Haag et al, J Mol Biol 243:537,1994). More may be discovered as the Human Genome Project advances the knowledge of our DNA sequence.
(Another interesting group of unitary pseudogenes are polymorphic sequences that are genes in some individuals and pseudogenes in others, with differing frequencies of pseudogenes in various populations. Examples include human genes for the chemokine receptor CCR5, for the alpha2-fucosyltransferase Se, for the cytochromes p450 2C19 and p450 2D6, for the enzyme thiopurine methyltransferase, and for the lipoprotein apo(a). For these examples, absence of the corresponding functional proteins is inconsequential most of the time, but can become clinically important -- whether beneficial or detrimental -- in certain environmental circumstances or in combination with other genes. In some cases the same gene product may be beneficial in some circumstances and detrimental in others, so that selection leads to an intermediate gene frequency.)
b. Classical duplicated pseudogenes
A much larger class of pseudogenes apparently arises from mishaps in a pattern of gene alteration that has been important in the evolution of normal functional genes: the pattern of duplication and differentiation (Ohta, Genome 31:304,1989; Holland et al., Dev Suppl 36:125, 1994). This pattern is evident from the frequent observation (in DNA from a variety of species) of blocks of sequences that have apparently been duplicated so that two or more repeats of similar sequences appear side by side, i.e. in tandem (see box 1).
Presumably, immediately after duplication each gene copy had an identical sequence. (See Figure 3.) But as DNA sequences are copied from generation to generation, mutations can accumulate independently in the duplicated sequence copies, with several possible consequences.
|
|
|
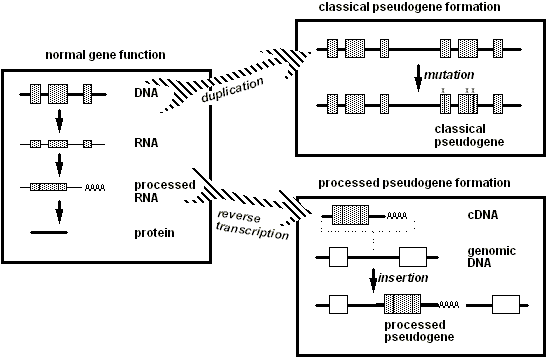
Figure 3. In normal gene function (left panel), DNA is transcribed into RNA, which is then "processed" by the removal of introns (the non-coding sequences between the gray boxes) and addition of a poly(A) tail. The mature processed RNA is then translated into a chain of amino acids to form a protein. The right panels illustrate the two pathways generating the classical duplicated pseudogene (top) and processed pseudogene (bottom). In the top pathway, DNA duplication generates two copies of the entire gene (upper right box), but mutations in one copy (represented by the "x"s) render it a pseudogene. In the other pathway a processed RNA transcript of a gene can become reverse transcribed into a cDNA copy (lower right box) that inserts back into cell's DNA at a random position in the genome, usually--as shown here--in the spacer DNA between genes (white boxes in the Figure). |
i. Some mutations may have no effect on the functioning of the gene.
| BOX1 |
|---|
|
Creationists commonly argue that features we observe in the DNA of modern species--presumably including tandemly repeated sequences--were designed specifically by an intelligent creator; whereas scientists view tandem repeat sequences as resulting from accidental DNA duplications. One argument in favor of the scientific view is that we can see examples of such genetic accidents occurring today in human DNA (as well as DNA from laboratory species) without apparent divine intervention. Because of ascertainment bias--that is, we find things only where we look for them--the best studied examples of duplications in humans are those that cause disease. One way DNA duplications can cause disease is if only part of a gene is duplicated, so that the resulting protein would have some amino acids repeated, thus altering the structure and function of the protein (Heikkinen et al., Am J Hum Genet 60:48, 1997; Hu and Worton Hum Mutat 1:3, 1992). When DNA duplications occur in somatic cells (i.e. outside the cell lineage contributing to egg and sperm) they cannot be passed on to future generations, but they can cause problems for the individual affected; for example, cases of cancer have been reported containing partial gene duplication in the cancer cells, while the duplication is absent from the normal body tissues (Schichman et al., Cancer Research 54: 4277, 1994), indicating that the duplication occurred during the life of the affected patient. Large duplications involving entire genes can create clinical problems if extra copies of an entire functional gene can produce harmful effects; while this is unusual, a well-studied example is the neurological disease Charcot-Marie-Tooth disease type 1A, in which an extra copy of the gene known as PMP-22 appears to be the culprit. Numerous cases of CMT1A have been reported where the duplication is present in the affected patient but not in either of the patient's parents, indicating that the duplication must have occurred in the germ cells of either parent or very early in the embryonic development of the patient (Eur Neurol 34:135, 1994). These examples make it clear that gene duplication is not just a hypothetical construct--invoked to explain tandem repeats that were created by inscrutable events in the distant past--but rather is a natural biochemical process that can be observed today in humans (it can also be studied in laboratory species as diverse as bacteria and fruit flies; Lupski et al. Am J Hum Genet 58:21, 1996). Genomes of modern vertebrate may reflect evidence of two ancient genome-wide duplication events that doubled the entire gene content (Sidow Curr Opin Genet & Devel 6:715,1996; Endo et al, Gene 205:19, 1997; Pebusque et al. Mol Biol Evol:1145, 1998); similar more recent doublings have been deduced in certain plants, frogs and even rats (Gallardo et al, Nature 401:341, 1999). |
ii. Other mutations may lead to a protein that has a
slightly different function from that of the original gene.
In fact, such differentiation of duplicated genes to
develop new functions in one copy apparently accounts for a
significant part of the expansion in complexity of the
genes of higher organisms. For example, the gene for a
primordial oxygen-carrying protein is thought to have
duplicated leading to separate genes encoding myoglobin
(the oxygen-carrying protein of muscle) and hemoglobin (the
oxygen-carrying protein of red blood cells). Then the
hemoglobin gene duplicated, and the copies differentiated
into the forms known as ![]() and
and
![]() . Later, both the
. Later, both the ![]() and
and ![]() hemoglobin genes duplicated several times
producing a cluster of hemoglobin-
hemoglobin genes duplicated several times
producing a cluster of hemoglobin-![]() -related sequences and a
cluster of hemoglobin-
-related sequences and a
cluster of hemoglobin-![]() -related sequences. The clusters include
functional genes that are slightly different, that are
expressed at different times during the development of the
embryo to the adult, and that encode proteins specifically
adapted to those developmental periods. The divergence
between the myoglobin and
-related sequences. The clusters include
functional genes that are slightly different, that are
expressed at different times during the development of the
embryo to the adult, and that encode proteins specifically
adapted to those developmental periods. The divergence
between the myoglobin and ![]() and
and
![]() genes occurred so long ago in
evolution that the shuffling of genetic information that
occasionally occurs in DNA has distributed these genes to
different chromosomes. The genes within the
genes occurred so long ago in
evolution that the shuffling of genetic information that
occasionally occurs in DNA has distributed these genes to
different chromosomes. The genes within the ![]() group and the
group and the ![]() group duplicated more recently in
evolution, and still lie in clusters.
group duplicated more recently in
evolution, and still lie in clusters.
iii. Finally, still other mutations that alter critical amino acids, that affect intron splicing or that create new stop codons, may completely destroy the function of a duplicated gene sequence and render it a pseudogene; indeed this is the fate of most gene duplicates (Lynch and Conery Science 290:1151, 2000). The kinds of mutations that destroy gene function again resemble those that have disabled crucial non-duplicated genes, thereby causing genetic diseases. Defective genes that are not duplicated tend to disappear from populations over time because individuals lacking a functional copy of the gene are less capable of surviving to produce offspring (unless the gene is no longer needed, as in unitary pseudogenes). However, when a defective gene exists alongside a normal duplicate copy, the continued function of the normal gene generally compensates for any mutations in the defective copy; the defective sequence is usually harmless and may be perpetuated in the DNA as a "classical duplicated" pseudogene. In general, each pseudogene of this type contains sequence resembling the entire gene--including both regulatory sequences lying outside the amino acid coding sequences, and introns that interrupt the coding sequences (see above). Numerous pseudogenes of this type have been found in DNA from a variety of organisms, including humans. For example, both the alpha and beta clusters of hemoglobin genes in humans include duplicated pseudogenes of this type.
Although most "classical" pseudogenes lie close to the gene from which they originated via tandem duplication, recently several laboratories have described a peculiar variety of duplicated pseudogenes located near the centromeres of several different chromosomes. (The centromeres are the chromosomal segments where--just before cell division--the two duplicated hot-dog-shaped chromosome copies appear tied together, as diagrammed in Figure 4, below). Apparently during primate evolution several DNA regions have undergone a poorly understood process which has distributed imperfect copies to the centromeric regions of multiple chromosomes. The genes in these copies include introns but are in many cases truncated and generally have multiple point mutations rendering them non-functional pseudogenes. Examples of these centromeric pseudogenes include sequences related to the adrenoleukodystrophy gene ALD (Eichler et al., Human Molec Genet 6:991, 1997), the creatine transporter gene SLC6A8 (Eichler et al. Human Molec Genet 5:899, 1996), the neurofibromatosis gene NF1 (Human Molec Genet 6:9, 1997) and a gene called FRG1 close to the facioscapulohumeral muscular dystrophy locus (Grewal et al., Gene 227:79, 1999).
c. Processed pseudogenes
An entirely different class of pseudogenes known as processed pseudogenes (see Figure 3, lower right panel) arises from naturally occurring insertions of extra gene copies derived from RNA transcripts. Three characteristics of these sequences suggest derivation from RNA:
i. Each processed pseudogene sequence resembles an RNA transcript in that the pseudogene's similarity to its "source gene" extends from the RNA initiation site to the RNA termination site, but does not include sequences that lie just outside the transcribed region, including regulatory sequences like promoters.
ii. These pseudogenes lack intron sequences that are normally transcribed into RNA but are then spliced out of the RNA before it is used to specify the amino acid sequence of a protein.
iii. They generally include the poly(A) "tail" characteristic of RNA transcripts that encode proteins.
Moreover, unlike the classical duplicated pseudogenes, which are usually found close to the functional gene from which they derived by duplication, processed pseudogenes are apparently inserted into DNA at random locations. This randomness is what one would expect from an RNA molecule that can float freely away from its source gene (from which it was originally transcribed) before a copy is reinserted back into the DNA. Even if it encodes a correct amino acid sequence, a processed pseudogene is usually non-functional because it lacks the regulatory sequences (like a transcriptional promoter) necessary for gene expression; as a non-functional extra copy such a sequence can accumulate random mutations under no selective pressure, i.e., without reducing the reproductive success of an organism that carries such mutations.
Processed pseudogenes should not be confused with a small number of retroposed gene copies that are actually functional genes. These can arise because, rarely, a DNA copy of a processed RNA may insert into DNA in such a way that the inserted copy can be actively expressed. In these cases, the DNA sequence may retain function and thus remain under selective pressure against accumulating mutations. Although such sequences--which can be called processed genes or retroposed genes--represent a tiny fraction of retroposed gene copies observed, more than a dozen have been discovered, especially as gene copies that are expressed specifically in the testis (Kleene et al, J Molec Evol 47: 275, 1998). These retroposed genes are easily distinguished from processed pseudogenes by their lack of crippling mutations. The fact that these few retroposed DNA copies are useful genes does not hint at any function for the much more numerous processed pseudogenes with multiple crippling mutations like stop codons that would preclude their expression as functional proteins.
Evolutionists as early as Darwin pointed to vestigial structures--such as the functionless eyes of blind cave-dwelling animals or the rudimentary pelvic bones of some snakes--as supporting the evolutionary viewpoint. These structures serve no apparent function that could explain their design by a creator, but they can easily be understood in the evolutionary perspective as deriving from structures that were functional in ancestral species. Vestigial genetic sequences--that is, pseudogenes--provide exquisite examples of vestigial structures, and thus especially compelling evidence for evolution. Such sequences can be studied in a variety of species; their relationship to their functional counterpart is obvious and quantitative (based on the number of sequence discrepancies between gene and pseudogene); and the subset of processed pseudogene can--with rare and easily recognizable exceptions--be assumed to have been totally functionless since the time of their origin. Finally, pseudogenes are a rich source of data because they are abundant. The recently completed sequence of human chromosome 22 (Dunham et al, Nature 402:489, 1999) identified 134 pseudogenes along with 545 genes in the sequenced region, which corresponds to about 1.1% of the human genome. If this sample is representative, we can expect roughly 15000 pseudogenes in the human genome.
2.2.2 Retroposons
How might a processed RNA sequence make its way back into DNA? In fact, processed pseudogenes are members of a much larger class of sequences known as retroposed elements, which I call here "retroposons," and which all involve RNA sequences that have been inserted into DNA. As discussed above, in normal cellular molecular physiology, genetic information passes from DNA to RNA to protein. Sometimes, however, a genetic accident occurs and the RNA gets reverse-transcribed into DNA ("retro" or backwards from the normal direction) and the DNA gets deposited back (or "retroposed") at some random position in the cell's DNA. (See Figure 4.) Some readers may recognize retroposition as the mechanism by which retroviruses like HIV--the AIDS virus--hide in the DNA of T cells of AIDS patients. As in the case of HIV, a critical requirement for all retroposition is activity of an enzyme called reverse transcriptase (RT), which generates a DNA copy of the RNA sequence. No gene encoding this enzyme is known to be present in the human genome other than in copies associated with elements that undergo retroposition. The RT enzyme has no known function for normal cellular physiology, although it appears that a specialized variant of reverse transcriptase is involved in maintenance of telomeres, repetitive sequences at the tips of chromosomes. Once a DNA copy of an RNA has been synthesized by RT, this DNA may be inserted into breaks in DNA that occur from time to time in the cell and that are normally sealed by a complex machinery of DNA repair enzymes required by all cells. Such breaks often occur at slightly different positions in the two DNA strands, producing "staggered ends"; retroposon sequences inserted between such ends are frequently flanked on both sides by short identical sequences created by repair of the two staggered ends.
|
|
|
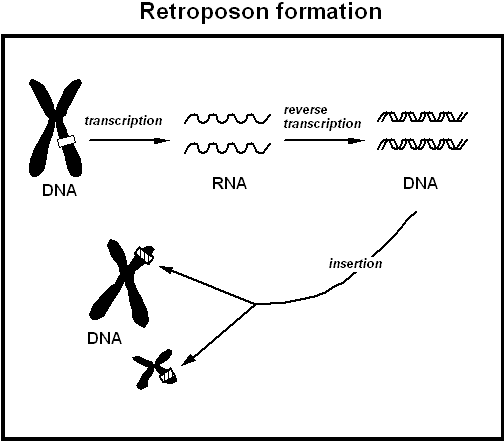
Figure 4. Retroposon formation occurs when a DNA sequence becomes transcribed into RNA, which then is "reverse transcribed" back into DNA. At the top left the Figure shows a chromosome as it appears just before cell division, looking like two hot dogs tied together at the centromere. One gene on a given chromosome can give rise to several pseudogenes, which generally insert randomly into different chromosomes. A similar mechanism spreads LINEs and SINEs, including Alu insertions. |
Of the many type of retroposons known to molecular biologists, I will mention four major classes found in human DNA. (For a recent review, see Prak and Kazazian, Nature Reviews, 1:134, 2000.)
a. Processed pseudogenes. In general, processed pseudogenes (described above) have been discovered as scientists have screened the genome (using techniques beyond the scope of this article) for sequences similar to known genes. Such screens turned up mutated versions with the "processed" features described above, leading to their identification as retroposons. Like any retroposon, this class could have entered the germline DNA (i.e. the DNA of sexual cells that allow propagation to future generations) only if the germ cells contained two components: RNA transcripts of the gene and reverse transcriptase (RT) to copy it back into DNA. Let us consider these two components in turn. Many of the best-known proteins are found only in specific differentiated tissues and are not expressed elsewhere. For example, hemoglobin is produced only in blood cells and their precursors, and the visual pigment proteins are produced only in the eye. The genes for such tissue-specific proteins are almost never transcribed in germ cells, and so they only rarely contribute to processed pseudogenes. In contrast, all cells have certain "housekeeping" proteins necessary for basic metabolic functions; RNA transcripts encoding these proteins are present in the germ cells and frequently contribute to processed pseudogenes. The glyceraldehyde-3-phosphate dehydrogenase gene for example is a housekeeping gene represented by about 10 processed pseudogenes in human DNA (Ercolani et al, JBC 263:15335,1988). Because, as mentioned above, processed pseudogene sequences do not include the promoter sequences necessary for the initiation of RNA transcription, once they are inserted into DNA these pseudogenes generally are not transcribed and thus cannot themselves represent a source gene for future retroposons. (This limitation contrasts with features of other retroposon classes, as we shall see below.) The other component necessary for retroposition is reverse transcriptase. RT is not present in most normal tissues in measurable amounts, although it can be expressed if a cell is infected with a retrovirus carrying the gene for this enzyme. Germ cells, however, are one cell type in which RT activity can be found in the absence of infectious retroviruses. In these cells the enzyme apparently derives from other retroposon elements--to be discussed below--that carry functional RT genes within their sequence.
b. SINEs. The best characterized class of Short Interspersed Elements (SINEs) in primates are known as Alu sequences. These are approximately 300 bp long and do not encode any protein sequence. The recent DNA sequence analysis of the human genome found about 1.1 million Alus, comprising 10.6% of the DNA (Nature 409:860, 2001). Unlike processed pseudogenes, which generally are not transcribed, Alu sequences include a segment that can act as an internal transcriptional promoter; thus each Alu insertion can potentially be transcribed into RNA, serving as the source for a new insertion. This property may partially explain how these sequences have become so abundant in our genomes. However, current evidence suggests that only a very few Alu sequences are active sources of transcripts; perhaps transcription from most copies is inhibited by the chromosomal environment of the insertion (Englander and Howard, JBC 270:10091, 1995). Even if Alu RNA transcripts exist in some germ cells, they re-insert into the DNA only rarely because this step requires reverse transcriptase, which may not be present in the same cells where the Alu RNA is being transcribed.
c. LINEs. Long Interspersed Elements represent a family of related sequences that are present in about 868,000 copies in our DNA, comprising about 20% of our genome (Nature 409:860, 2001). They differ from the Alu sequences in being much longer--up to about 7000 basepairs--and in containing two potential coding sequences. One of these coding sequences bears similarity to active RT genes. Although in most LINE copies the RT gene contains numerous mutations that would prevent it from encoding any functional RT enzyme, certain LINE copies do encode active reverse transcriptase. Moreover, the regulatory regions just outside the coding sequences of the LINEs cause expression of the genes selectively in germline cells. LINEs thus have several properties expected of "selfish" DNA sequences that can spread in the host DNA simply because they encode their own machinery for spreading. The LINEs can be expressed in germline cells as RNA, and the rare copies that encode functional RT can enable the reverse transcription of the LINE RNA back into a DNA copy which can then insert into new locations in the DNA of the germline cell; when such a cell matures to egg or sperm, transmission of the new LINE to future generations can occur. Apparently the RT often falls off the RNA before reverse transcription is complete, since most LINE copies are truncated at their 5' ends. It is possible that the LINE-encoded reverse transcriptase activity can also produce the reverse-transcribed copies of other RNAs--such as Alu transcripts and RNA transcripts of genes--that lead to new insertions of Alu sequences and processed pseudogenes into cellular DNA.
d. Endogenous retroviruses. Infectious retroviruses were discovered as agents of human disease and have been intensively studied. They are the most complex of retroposing elements and may have evolved from simpler ones described above. All contain two identical non-coding Long Terminal Repeats (LTRs) at their ends as well as three genes known as gag, pol and env. These genes are encoded in the virus not by DNA but by RNA. The pol gene encodes reverse transcriptase, and may also encode additional enzymatic activities. The env gene encodes proteins that coat the outside surface (envelope) of the infectious virus. The gag gene encodes additional proteins necessary for processing the viral components. The structure common to all retroviruses is thus LTR-gag-pol-env-LTR. The "left" LTR includes regulatory sequences that can initiate RNA transcription towards the right, into the gag-pol-env-LTR; the "left" LTR is then recopied from the "right" LTR by a complex mechanism. Infectious retroviruses include HTLVI, which causes a kind of leukemia in humans, and HIV, which causes AIDS. These viruses typically infect specific kinds of white blood cells--lymphocytes--and insert reverse-transcribed copies of their RNA genes into the DNA of these cells. Soon after the discovery of infections retroviruses, scientists noticed that similar sequences were present in the DNA of many mammalian species, including humans; these copies are called endogenous retroviruses, and presumably represent the consequences of ancient retroviral infections of germline cells. In human DNA there are about 8 different classes of endogenous retroviruses with members of each class varying in number from one or two to more than 50 copies. Essentially all of these endogenous retroviruses contain mutations that would disrupt the function of their genes, as would be expected if they inserted millions of years ago with no selective pressure to maintain the function of the genes. In addition, the duplicated LTR sequences represent potential targets for "homologous recombination" events that delete the DNA between the corresponding region of the LTRs, leaving only a single composite LTR sequence; many more copies of these isolated LTR fragments exist in the DNA than complete retroviral copies.
3. How ancient errors can persist in modern species
How could each of the several kinds of non-functional sequences mentioned above, arising in a germline cell of a particular individual, come to be preserved in all individuals of a species? One possibility is that each of these sequences happened to lie close to an advantageous gene that became prevalent in a population by natural selection (the pseudogene or retroposon "rode on the coat-tails" of the nearby advantageous gene; see Nurminsky et al, Nature 396:572,1998; Nurminsky et al. Science 291:128, 2001). Possibly such non-functional sequences arise at a high frequency and we see only those few that are preserved by such indirect influences or by chance events in small populations.
The extra burden of carrying along even a large pseudogene sequence--for example, 100,000 nucleotides--is insignificant for a mammalian cell with approximately three billion nucleotides' worth of information. In any case, there is no known "proofreading" mechanism by which the cell might distinguish non-functional from functional DNA and selectively eliminate what it does not need. Functionless DNA sequences that scientists have inserted into the DNA of mice or other species are faithfully passed to descendants, and naturally occurring pseudogenes and retroposons apparently behave similarly. The accumulation of functionless DNA is not completely unopposed; deletions of DNA do occur, but apparently as rare accidents that do not discriminate between functional and non-functional sequences. Deletions that remove crucial functional genes have been recognized as rare causes of genetic diseases; decreased fitness of the individuals that carried them would tend to eliminate DNA copies with such deletions. Other deletions that by chance do not remove any functional genes could eliminate some useless DNA including pseudogenes and retroposons; but an individual with such a deletion would have no particular selective advantage as a result of the deletion, so spread of DNA copies carrying the deletion into the population at large would be no more likely than the spread of any other inconsequential mutation. Thus such deletion events are clearly an inefficient "garbage removal" mechanism; and, as an inevitable consequence of this inefficiency, substantial amounts of functionless "garbage" sequences have accumulated between the functional genes of mammals. This is a characteristic of the genetic material that was not appreciated until recombinant DNA technology enabled molecular biologists to look beyond amino acid sequences to the structure of DNA itself. Although the high content of "junk DNA" was initially surprising when it was discovered, our current understanding of the mechanisms of genome expansion (duplication and insertion) and the apparent lack of significant selective pressure to minimize genome size combine to make the accumulation of useless sequences in our DNA seem inevitable.
4. The argument from DNA to evolution: Shared pseudogenes and retroposons
The crucial observation relating the discovery of pseudogenes and retroposons to the theory of evolution is this: some pseudogenes and retroposons are shared between different species, as though they were copied from a pseudogene or retroposon in a common ancestor. Let's examine examples from each of the classes of "errors" we have discussed above.
| BOX 2 |
|---|
|
The shared galactosyltransferase pseudogenes are fascinating for a reason that complicates their use in arguing against creationists: evidence suggests that there may have been a selective advantage to mutations that inactivated this gene. The enzyme product of the gene catalyzes the production of a particular carbohydrate molecule that is found on cell membranes of mammals who possess the enzyme, but also on certain infectious bacteria. Individuals infected with such bacteria would benefit from mounting an immune attack on this carbohydrate molecule, but if the same carbohydrate appeared on their own cells such an attack could damage their own tissues. Therefore, individuals who carry mutations in the enzyme--and thus would not make the carbohydrate on their own cells--would be free to mount an immune attack focused on this molecule, protecting them against many bacteria without danger of damaging their own tissues. Therefore, selective pressure would have led to spread of gene copies that had undergone crippling mutations. Creationists could reasonably argue that such mutations could have occurred independently in different species as examples of recent microevolution after independent creation of the species. It is possible that different mutations did inactivate the gene independently in several primate ancestors. However, the human and chimpanzee galactosyltransferase pseudogenes have identical crippling mutations; therefore, it is most likely that the gene was inactivated in a common human/chimp ancestor. |
4.1. Shared unitary pseudogenes. Many of the unitary pseudogenes in humans described previously are shared with other primates. By "shared" I mean more than simply that the same gene is inactive in two different species, since that situation could result if the corresponding genes of the two species were inactivated separately by independent mutations. Instead, in all the examples I describe, the pseudogenes in primates carry many of the same crippling mutations found in the corresponding human pseudogenes. Since independent random mutations would not be likely to be identical in two different species, the identically mutated pseudogenes are strong evidence that the mutations occurred in a common ancestral species.
For the example of the GLO unitary pseudogene of humans, it is known that vitamin C is required in the diet of other primates, (though not for other mammals except guinea pigs). The theory of evolution would make the strong prediction that primates should also be found to have GLO pseudogenes and that these would carry similar crippling mutations to the ones found in the human pseudogene. This prediction was stated in earlier versions of the present essay. A test of this prediction has recently been reported. A small section of the GLO pseudogene sequence was recently compared from human, chimpanzee, macaque and orangutan; all four pseudogenes were found to share a common crippling single nucleotide deletion that would cause the remainder of the protein to be translated in the wrong triplet reading frame (Ohta and Nishikimi BBA 1472:408, 1999).
The RT6 gene mentioned above (2.2.1.a) encodes a protein of about 230 amino acids expressed on the surface membrane of T lymphocytes of rodents; both the human pseudogene and its chimpanzee homolog contain mutations producing the same three stop codons that would prevent the synthesis of an RT6 protein (Haag et al, M Mol Biol 243:537,1994). Several of the human odorant receptor pseudogenes mentioned above are found in other primates, and share the same defects as the human pseudogenes (Rouquier S et al., Nat Genet18:243,1998; Rouquier S, et al. Human Molec Genet &:1337,1998;Sharon et al., Genomics 61:24,1999). The human NPY1 receptor pseudogene shares a critical frameshift mutation with primate homologs (Matsumoto et al., J Biol Chem 271:27217, 1996). The human urate oxidase pseudogene shares three crippling mutations with the chimpanzee and orangutan pseudogenes (Wu et al, J Mol Evol 34:78, 1992). In addition, the galactosyltransferase pseudogene present in the human genome is shared with apes and Old World monkeys ( Galili and Swanson, PNAS 88:7401, 1991) although the evolutionary interpretation of these shared galactosyltransferase pseudogenes is complex because there may have been selective pressure to inactivate this enzyme (see Box 2).
In summary, although unitary pseudogenes are relatively rare in humans, most of the reported examples are shared with other non-human primates.
(The only other examples of human unitary pseudogenes I know of are unique to humans, apparently having acquired their crippling defects after the human-chimpanzee split; they are therefore of interest as potentially contributing to the physiologic differences between these two species. These pseudogenes correspond to a type I hair keratin [Hum Genet 108:37, 2001], CMP-sialic acid hydroxylase [Chou et al., PNAS 95:11751, 1998]. flavin-containing monooxygenase-2 (FMO2) [J Biol Chem 273:30599, 1998], CMP- N-acetylneuraminic acid hydroxylase [Hayakawa et al.,PNAS 98:11399, 2001] and the V10 variable gene of the human T-cell receptor gamma locus [Zhang et al., Immunogenetics 43:196, 1996]. Readers are invited to let me know about additional unitary pseudogenes.)
4.2. Classic duplicated pseudogenes. There are many examples of shared pseudogenes of this type; I will describe only one. The steroid 21-hydroxylase gene encodes an enzyme involved in metabolism of steroid hormones. In human DNA, the 21-hydroxylase gene sequence, as well as an adjacent gene encoding "complement C4," has been duplicated; i.e., nearly identical copies of DNA segments lie adjacent to each other, each copy containing a complement C4 gene and a steroid 21-hydroxylase sequence. However, only the "B" copy of the 21-hydroxylase gene is functional; the "A" copy in all humans is a pseudogene, i.e., it contains multiple mutations including an 8 bp deletion that would prevent its function. The corresponding "A" copy sequence of chimpanzee has been examined; it contains the same crippling 8 bp deletion seen in the human pseudogene (Kawaguchi, Am J Hum Genet 50:766-80, 1992).
Many of the peculiar centromeric pseudogenes described above (in section 2.2.1.b) are also conserved in other primates (Eichler et al., Human Molec Genet 5:899, 1996; Regnier et al, Human Molec Genet 6:9, 1997; Grewal et al., Gene 227: 79, 1999).
4.3. Processed pseudogenes. Because human DNA may contain roughly four times more processed pseudogenes than classic duplicated pseudogenes (extrapolating from data from chromosome 22 [Dunham et al, Nature 402:489, 1999]), there are many more examples of processed pseudogenes (than classical pseudogenes) shared between species. I will describe one that my colleagues and I discovered: a pseudogene derived from the gene encoding epsilon immunoglobulin--a kind of antibody that participates in allergic reactions. In our studies aimed at investigating the basis for allergy we discovered a sequence that resembled the epsilon immunoglobulin gene except that it had no introns, it had multiple crippling mutations, it had on its end a sequence of almost continuous "A"s (looking like a slightly mutated poly(A) tract), and it was located on a different chromosome (chromosome 9) from that of the functional gene (chromosome 14) (Max et al. Cell 29:691, 1982; Battey et al. PNAS 79:5956, 1982). Our evidence suggested that this processed pseudogene also existed in chimpanzee DNA, and subsequent detailed investigations from other laboratories (Kawmura and Ueda, Genomics 13:194,1992) demonstrated nearly identical pseudogenes exist in chimpanzee, gorilla, orangutan and Old World monkeys. As in the case of all DNA insertions shared by different species (see other examples below), the argument that these sequences were not created independently but descended from a common ancestral insertion is bolstered by the demonstration that the insertions occurred in the same position in the DNA of each species, i.e., the DNA that surrounds the insertion is very similar between species--as close to identical as might be expected given the occurrence of mutations that are not selected against.
4.4. SINEs. Of the roughly one million copies of
Alu sequences in the human genome, only a small fraction
have been compared between human and other primate species.
However, in several long segments of DNA where the
corresponding sequences have been obtained in human and
chimpanzee DNA, almost all of the Alu sequences are shared
between these two species. For example, in the cluster of
![]() -globin genes referred to
above, all seven of the Alu sequences found in human DNA
are present in chimpanzee, embedded in exactly the same
positions (Sawada et al. J Mol Evol 22:316, 1985). The same
is true of the seven Alu sequences near a pseudogene
derived from the single-copy cdc27hs gene (Gonzalez et al.,
Genomics 18:29, 1993).
-globin genes referred to
above, all seven of the Alu sequences found in human DNA
are present in chimpanzee, embedded in exactly the same
positions (Sawada et al. J Mol Evol 22:316, 1985). The same
is true of the seven Alu sequences near a pseudogene
derived from the single-copy cdc27hs gene (Gonzalez et al.,
Genomics 18:29, 1993).
The sequences of many Alu repeats in human DNA have been compared, allowing classification into several families, based on the degree of sequence similarity. Members of certain families are found in DNA of many diverse primates, whereas other families appear to have been dispersed more recently as they are not shared by other species. Several examples of insertions of the "youngest" family are known to be polymorphic in the human population: i.e., they occur in some individuals but not others. Indeed, the frequency of certain Alu insertions in different human populations has been used to deduce likely patterns of migration and gene mixing in our human ancestors. Such observations are consistent with the insertion of such Alu copies after the evolution of humans. Further, the excellent health of individuals who lack particular Alu insertions supports the view that these insertions do not serve any important function in human physiology.
4.5. LINEs. Numerous LINE sequences have been found at the same position in the DNA of humans and other species, including examples in the globin locus, visual pigment genes, and intestinal alkaline phosphatase (reviewed by Smit et al. J Mol Biol 246:401,1995). Some of the reported examples are shared by species as disparate as human and cow, indicating insertions in very early mammalian ancestors.
4.6. Endogenous retroviruses. Because endogenous retroviruses are less numerous than the other nonfunctional DNA sequences discussed here, and because a relatively tiny fraction of the known human DNA sequences have been compared between species, there is a dearth of examples of shared endogenous retroviruses. However, at least five different examples of nearly identical retroviral sequences embedded at the same position in human and chimpanzee DNA have been reported (Bonner et al. PNAS 79:4709, 1982; Dangel et al. Immunogenetics 42:41, 1995; Svensson et al Immunogenetics 41:74,1995; Medstrand & Mager J Virol 72:9782, 1998; Barbulescu et al. Curr Biol 9:861, 1999), all apparently examples of retroviruses that were "caught" by ancestors of ours millions of years ago. One can anticipate that additional examples will be discovered as more sequence data become available, especially from the Y chromosome, which has been described as a "graveyard" for endogenous retrovirus sequences for both human and chimpanzee (Kjellman et al. Gene 161:163, 1995).
4.7 Implications of functionless sequences shared between species
All of the examples of functionless sequences shared between humans and chimpanzees reinforce the argument for evolution that would be compelling even if only one example were known. This argument can be understood by analogy with the legal cases discussed earlier in which shared errors were recognized as proof of copying. The appearance of the same "error"--that is, the same useless pseudogene or Alu sequence or endogenous retrovirus at the same position in human and ape DNA--cannot logically be explained by independent origins of the two sequences. The creationist argument discussed earlier--that similarities in DNA sequence simply reflect the creator's plans for similar protein function in similar species--does not apply to sequences that do not have any function for the organism that harbors them. The possibility of identical genetic accidents creating the same two pseudogene or Alu or endogenous retrovirus independently in two different species by chance is so unlikely that it can be dismissed. As in the copyright cases discussed earlier, such shared "errors" indicate that copying of some sort must have occurred. Since there is no known mechanism by which sequences from modern apes could be copied into the same position of human DNA or vice versa, the existence of shared pseudogenes or retroposons leads to the logical conclusion that both the human and ape sequences were copied from ancestral sequences that must have arisen in a common ancestor of humans and apes.
This evidence for a common ancestor clinches the argument for human/ape evolution that follows from shared functionless sequences. Although the most numerous documented examples of such sequences shared between different species happen to link humans and apes (see for example Hamdi et al, J Mol Biol 284:861, 1999), this simply reflects the fact that the DNA of humans has been studied more intensively than DNA from any other higher species, while considerable homologous chimpanzee sequence is also known. It is obvious, however, that the identical logic could be used to link other species on different branches of the evolutionary tree, and such examples have been reported, e.g. SINEs clarifying relationships between rodent species (Furano J Biol Chem. 270: 25301, 1995; Verneau et al, PNAS 95: 11284, 1998) or linking horses to rhinoceros (Gallagher et al, Mamm Genome:140, 1999) or establishing the phylogenetic affiliations of tarsiers (Schmitz et al., Genetics 157:777, 2001) or pecoran ruminants (Nijman et al J Mol evol 54:9, 2002). Species as disparate as humans and mice have been linked by examples of the ancient SINE family known as MIRs (Mammalian-wide Interspersed Repeats; see Smit and Riggs, Nucleic Acids Research 23:98, 1995; Jurka et al, Nucleic Acids Research 23:170, 1995) that were found embedded at the homologous location in the human and murine myoglobin and N-myc genes (Donehower, Nucleic Acids Research 17:699, 1989; note that at the time of this description the conserved sequence was not recognized as a SINE). Additionally, ancient LINE insertions link humans to cow, as mentioned above (similar LINE inserts lying upstream of the intestinal alkaline phosphatase genes in both species), as well as to rat (similar LINE insertions in the first intron of the alpha2 subunit of the sodium-potassium ATPase genes; Smit et al, J Mol Biol 246:401, 1995) and to mouse (e.g. LINE insertions in the mnd2 region of chromosome 2p13 [Jang et al, Genome Res 9:51, 1998] and near the CD4 gene at human chromosome 12p13 [Ansari-Lari et al Genome Res 8:29, 1998]). With additional sequence comparisons of long homologous stretches of human and mouse DNA anticipated from the Human Genome Project and Mouse Genome Project, additional LINE sequences shared between these species will likely be discovered.
A particularly impressive example of shared retroposons has recently been reported linking cetaceans (whales, dolphins and porpoises) to ruminants and hippopotamuses, and it is instructive to consider this example in some detail. Cetaceans are sea-living animals that bear important similarities to land-living mammals; in particular, the females have mammary glands and nurse their young. Scientists studying mammalian anatomy and physiology have demonstrated greatest similarities between cetaceans and the mammalian group known as artiodactyls (even-toed ungulates) including cows, sheep, camels and pigs. These observations have led to the evolutionist view that whales evolved from a four-legged artiodactyl ancestor that lived on land. Creationists have capitalized on the obvious differences between the familiar artiodactyls and whales, and have ridiculed the idea that whales could have had four-legged land-living ancestors. Creationists who claim that cetaceans did not arise from four-legged land mammals must ignore or somehow dismiss the fossil evidence of apparent whale ancestors looking exactly like one would predict for transitional species between land mammals and whales--with diminutive legs and with ear structures intermediate between those of modern artiodactyls and cetaceans (Nature 368:844,1994; Science 263: 210, 1994). (A discussion of fossil ancestral whale species with references may be found at http://www.talkorigins.org/faqs/faq-transitional/part2b.html#ceta) Creationists must also ignore or dismiss the evidence showing the great similarity between cetacean and artiodactyl gene sequences (Molecular Biology & Evolution 11:357, 1994; ibid 13: 954, 1996; Gatesy et al, Systematic Biology 48:6, 1999).
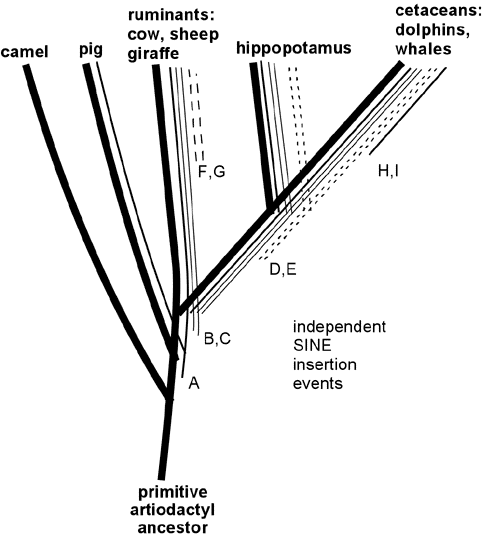
Recently retroposon evidence has solidified the evolutionary relationship between whales and artiodactyls. Shimamura et al. (Nature 388:666, 1997; Mol Biol Evol 16: 1046, 1999; see also Lum et al., Mol Biol Evol 17:1417, 2000; Nikaido and Okada, Mamm Genome 11:1123, 2000) studied SINE sequences that are highly reduplicated in the DNA of all cetacean species examined. These SINES were also found to be present in the DNA of ruminants (including cows and sheep) but not in DNA of camels and pigs or more distantly related mammals such as horse, elephant, cat, human or kangaroo. These SINES apparently originated in a specific branch of ancestral artiodactyls after this branch diverged from camels, pigs and other mammals, but before the divergence of the lines leading to modern cetaceans, hippopotamus and ruminants. (See Figure 5.) In support of this scenario, Shimamura et al. identified two specific insertions of these SINES in whale DNA (insertions B and C in Figure 5) and showed that in DNA of hippopotamus, cow and sheep these same two sites contained the SINES; but in camel and pig DNA the same sites were "empty" of insertions. More recently, hippopotamus has been identified as the closest living terrestrial relative of cetaceans since hippos and whales share retroposon insertions (illustrated by D and E in Figure 5) that are not found in any other artiodactyls (Nikaido et al, PNAS 96:10261, 1999). The close hippo-whale relationship is consistent with previously reported sequence similarity comparisons (Gatesy, Mol Biol Evol 14:537, 1997) and with recent fossil finds (Gingerich et al., Science 293:2239, 2001; Thewissen et al., Nature 413:277, 2001) that resolve earlier paleontological conflicts with the close whale-hippo relationship. (Some readers have wondered: if ruminants are more closely related to whales than to pigs and camels, why are ruminants anatomically more similar to pigs and camels than they are to whales? Apparently this results from the fact that ruminants, pigs and camels changed relatively little since their last common ancestor, while the cetacean lineage changed dramatically in adapting to an aquatic lifestyle, thereby obliterating many of the features -- like hooves, fur and hind legs -- that are shared between its close ruminant relatives and the more distantly related pigs and camels. This scenario illustrates the fact that the rapid evolutionary development of adaptations to a new niche can occur through key functional mutations, leaving the bulk of the DNA relatively unchanged. The particularly close relationship between whales and hippos is consistent with several shared adaptations to aquatic life, including use of underwater vocalizations for communication and the absence of hair and sebaceous glands.) Thus, retroposon evidence strongly supports the derivation of whales from a common ancestor of hippopotamus and ruminants, consistent with the evolutionary interpretation of fossils and overall DNA sequence similarities. Indeed, the logic of the evidence from shared SINEs is so powerful that SINEs may be the best available characters for deducing species relatedness (Shedlock and Okada, Bioessays 22:148, 2000), even if they are not perfect (Myamoto, Curr. Biology 9:R816, 1999).
|
|
|
Figure 5. Specific SINE insertions can act as "tracers" that illuminate phylogenetic relationships. This figure summarizes some of the data on SINEs found in living artiodactyls and shows how the shared insertions can be interpreted in relation to evolutionary branching. A specific SINE insertion event ("A" in the Figure) apparently occurred in a primitive common ancestor of pigs, ruminants, hippopotamus and cetaceans, since this insertion is present in these modern descendants of that common ancestor; but it is absent in camels, which split off from the other species before this SINE inserted. More recent insertions B and C are present only in ruminants, hippopotamus and cetaceans. Insertions D and E are shared only by hippopotamus and cetaceans, thereby identifying hippopotamus as the closest living relative of cetaceans (at least among the species examined in these studies). SINE insertions F and G occurred in the ruminant lineage after it diverged from the other species; and insertions H and I occurred after divergence of the cetacean lineage. |
While some creationists accept the evidence for the natural selection of minor variants (e.g. the divergence of Darwin's finches on the Galapagos islands), which they call "microevolution," most creationists deny that evolution can explain more significant changes, which they designate "macroevolution." However, the shared pseudogenes/retroposons described here provide strong evidence that humans share common ancestors with species as disparate as monkeys, cows and mice. Thus, even though we may lack convincing evidence that any particular fossil is ancestral to a specific modern species, and even though we do not have fossil evidence that clearly identifies the last common ancestor between humans and cows or between whales and ruminants, we can be confident from the shared errors described here that these common ancestral species existed. This conclusion in turn implies that significant novel characteristics (e.g. human upright walking and brain development, and the cetacean adaptations to aquatic life) must have developed between the time the respective common ancestors lived and the present day. These changes are clearly extensive enough to be called "macroevolution," so the "argument from shared errors" is powerful evidence for macroevolution. This conclusion seems solid, since no alternative explanation of these shared errors consistent with independent origin of these animal species has been proposed in the scientific literature.
Clearly the "shared errors" argument provides strong evidence for macroevolutionary changes in the evolution of mammals, and therefore refutes a commonly held creationist position. But to be fair we should be clear that this argument does not buy the whole evolutionist ballgame. Although the evidence of shared errors implies common descent of diverse mammalian species, it does not address whether these species evolved from their last common ancestors through the Darwinian mechanisms of mutation and natural selection or through other alternative mechanisms. Another limitation is that there are no examples of "shared errors" that link mammals to other branches of the genealogic tree of life on earth. For example, although species as diverse as worms, yeast and plants have LINE elements in their genomes, no examples of specific LINE insertions at homologous positions between any mammal and non-mammal have been reported to my knowledge (though I welcome input on this point from readers). Such examples might be expected to be hard to find, since the last common ancestors of mammals and reptiles are thought to have lived more than 200 million years ago, long enough that sequence similarities that once existed in functionless DNA like pseudogenes and retroposons may have been largely obliterated by the accumulation of numerous mutations. Therefore, the evolutionary relationships between distant branches on the evolutionary genealogic tree must rest on other evidence besides "shared errors." (Such evidence might include other "rare genomic changes" (RGCs) besides retroposon insertion, such as insertion or deletion of coding sequences or introns (e.g. Venkatesh et al., PNAS 98:11382,2001), chromosomal translocations and inversions revealed by comparative cytogenetics, and variants in the genetic code, all summarized in Rokas and Holland, Trends Ecol & Evol 15:454, 2000. Species relatedness can also be inferred from traditional sequence similarity trees based comparisons of the corresponding genes from different species.) As a final and rather obvious limitation of the "shared errors" argument, it should be clear that this argument does not bear on origin-of-life issues, which creationists commonly lump with evolution.
5. Creationists' responses to the argument from shared functionless sequences
Creationists tend to avoid mentioning the argument presented in this essay since it provides persuasive evidence for evolution, but creationist spokesman Duane Gish has commented on the argument when he has been confronted with it in debates; and a few other creationist discussions of pseudogenes have appeared. Let us first examine several of Dr. Gish's responses.
5.1 Some processed "pseudogenes" are functional, so they could be examples of "similar design for similar function."
As mentioned above (2.2.1.c), reverse-transcribed copies of RNA transcripts of genes may, rarely, insert into the DNA near an existing promoter or in some other way that allows their transcription in a manner that is useful for the organism. Such copies (which are really processed genes rather than processed pseudogenes) may therefore provide some function that provides selective pressure against crippling mutations. Several examples of this possibility have been reported, as mentioned above in section 2.2.1.c; and these could be interpreted as "similar design for similar function." But these examples share a feature that clearly distinguishes them from the hundreds of examples of useless processed pseudogenes reported: they lack crippling mutations that would preclude function, and thus remain capable of encoding a useful protein. Among bone fide processed pseudogenes--i.e. retroposed gene copies with multiple crippling mutations such as stop codons--no examples with documented function have been reported [See note added below] . (Readers who believe that there are examples contradicting this statement are invited to contact me with the literature references; I will modify this article as necessary.) Thus Dr. Gish's argument simply reflects his erroneous lumping together of two distinct classes of retroposed gene copies: processed genes and processed pseudogenes. And Dr. Gish has not yet offered any argument that would explain--in terms of intelligently designed function--the numerous examples of shared retroposed sequences that, unlike pseudogenes, do not even derive from DNA that has a functional role.
Note added May 5, 2003: An example of a pseudogene with documented function has just been reported. A sequence called Makorin1-p1 does not encode a functional protein, but its mRNA transcript stabilizes the mRNA of the functional Makorin1 gene from which it was apparently derived (Hirotsune et al, Nature 423:91,2003). Presently available knowledge from the published literature does not suggest functional roles for processed pseudogenes other than Makorin1-p1. It is possible that future studies will discover some other pseudogenes with apparent function (but see the next section).
5.2 Some organs previously thought to be vestigial have more recently been found to have function; we know too little about these newly discovered DNA features to be confident that function will not be discovered for them in the future.
Imagine a defendant at a murder trial defending himself--against overwhelming incriminating evidence--with the parallel argument: that since some convicted criminals have later been exonerated, he (the current defendant) should therefore be acquitted now, because someday in the future, evidence might be found to clear him! This defense would be as ridiculous as Dr. Gish's argument is. Scientists (and juries) must draw their conclusions based on the best evidence available at the time. It is true that later evidence may exonerate a convicted criminal or overturn a scientific theory. This possibility should foster humility and caution us against dogmatic conclusions (and perhaps against the death penalty); but it should not dissuade us from drawing the most reasonable conclusions from the data at hand. Our present knowledge supports the interpretation that most shared pseudogenes/retroposons are evidence for common descent and macroevolution. If in the future--for a particular Alu or LINE-1 or endogenous retrovirus sequence that is shared between humans and another species--evidence of function is discovered, then this particular sequence could indeed be reasonably interpreted by the creationist paradigm of "similar sequence designed for similar function"; and so this retroposon would have to be removed from list of shared functionless sequences that provide evidence for evolution. The hundreds of thousands of remaining examples on this list would continue to offer valid support for evolution.
Furthermore, while these vestigial DNA sequences were discovered more recently than the vestigial organs known in Darwin's time, we know enough about how they arise that we do not need to postulate any mysterious designer or unknown function to explain them. We know that the prerequisites for the formation of SINEs and other retroposons--i.e., RNA transcripts and reverse transcriptase--are present at low levels in germline cells studied in the laboratory, where they would be able, without any supernatural intervention, to generate retroposons that could be transmitted to future generations. This fact would predict that retroposon insertions must be occurring at some frequency even today. Indeed, specific insertions of Alu sequences into DNA of living individuals have been documented. For example, an Alu element was found inserted into the DNA of a patient with neurofibromatosis I, damaging the gene associated with this disease (Wallace et al. Nature 353:6347, 1991). The patient's father and mother had intact gene copies with no Alu insertion, so the insertion must have occurred in the germ cells of either parent or very early in the embryonic development of the patient. Similarly, a freshly inserted LINE element was found to have damaged the gene for a blood clotting protein, causing hemophilia in another patient whose parents both lacked this insertion (Kazazian et al. Nature 332:164, 1988). (Other examples of LINE or Alu insertion causing diseases are reviewed by Kazazian [in Curr Opin Genet & Devel 8:343, 1998] by Miki [Human Genetics 43:77, 1998] and by Deininger and Batzer [Molec Genet & Metab 6:183, 1999].) New retroposition events are estimated to occur in from 1% to 10% of the human population (Kazazian Nature Genet 22:130, 1999). Carlton et al (Mamm Genome 6:90, 1995) observed de novo appearance of a processed pseudogene when they provided a source of reverse transcriptase by infecting cultured cells with a retrovirus; while Esnault et al. (Nature Genet 24:363, 2000) and Wei et al. (Molec Cell Biol 21:1439, 2001) observed processed pseudogene formation resulting from the RT of a human LINE element. Using a sensitive assay for detecting retroposition, Maestre et al. (EMBO J 14:6388, 1995) were able to detect retroposed copies of a marked gene sequence being inserted into the DNA of human cells as they were growing in the laboratory even without the addition of exogenous reverse transcriptase. Furthermore, Jensen and Heidmann (EMBO J 10:1927,1991) detected ongoing retroposition of a marked LINE copy in Drosophila.
Recently Feng et al (Cell 87: 905, 1996) demonstrated that the active reverse transcriptase enzyme encoded by a freshly inserted LINE copy has an additional unexpected activity: it is an endonuclease--that is, it is able to cause nicks in DNA that could serve as insertion points for new retroposition events. In fact, this endonuclease cuts DNA with particular sequence characteristics, and the same characteristics were observed in the insertion positions of several randomly selected LINE copies from human DNA. (See also Cost and Boeke, Biochemistry 37:18081, 1998). This result suggests that LINE sequences are so well adapted for "selfish" replication in the genome that they do not depend on randomly generated breaks in DNA for their insertions, but generate their own cuts. To test this idea, Moran et al. (Cell 87:917, 1996; see also Ostertag et al, Nucl Ac Res 28:1418, 2000) constructed a LINE sequence designed so that if it generated any new retroposed copies in any cells, these cells could be selected and counted. When this sequence was put into human tissue culture cells, newly retroposed copies were routinely produced. By testing the effects of mutations in various segments of the LINE sequence, it was shown that efficient retroposition required both the reverse transcriptase activity and the endonuclease activity present in the same protein. (This protein is also required for efficient LINE-induced processed pseudogene formation [Esnault et al. Nature Genet 24:363, 2000]).
Observations like these reinforce the notion that the retroposon sequences we observe in our DNA and the DNA of other mammals were not created by mysterious forces acting only in the ancient past for inscrutable purposes, but by simple genetic accidents that occur at low frequency as a result of quirks of cellular biochemistry, and which serve no purpose. The fact that a very few of these genetic accidents may create some beneficial function (Britten RJ PNAS 93:9374,1996; Britten RJ Gene 205:177,1997) does not weaken this interpretation at all; such events are simply examples of rare beneficial mutations whose occurrence forms the basis for adaptive evolutionary change and whose existence seems so difficult for the creationists to swallow. As is the case for most mutations, the overwhelming majority of retroposon insertions occur in the non-functional DNA between genes, and have no effect on the cell or organism; and it is this vast set of insertions, shared between species, that provide the basis for the present argument supporting evolution.
5.3 If all these sequences were really nonfunctional, they would have been eliminated over evolutionary time.
This argument reflects ignorance of the facts discussed above in section 3. To repeat: no mechanism is known by which non-functional DNA sequences might be distinguished from functional ones and targeted for elimination by cellular enzymes. Bacteria do appear to be under selective pressure to eliminate nonfunctional DNA; bacterial chromosomes have very little DNA between genes, perhaps because competition under conditions of rapid growth may favor chromosomes that replicate quickly--i.e. short ones--and therefore may select for cells that have deleted any non-functional DNA. But there is no evidence for such selective pressure for mammalian chromosomes, in which genes are widely separated from one another and in which nonfunctional regions apparently constitute 90-95% of the DNA. Indeed, one might ask: why then are our chromosomes not stuffed with retroposon sequences at an even higher frequency than actually observed? A reasonable answer is that our ancestors were under selective pressure to suppress retroposition, since high frequencies of retroposon insertion would increase the rate of genetic damage caused by crippling insertions into genes. Furthermore, it is conceivable that a larger fraction of our DNA originated through retroposition than we can now recognize; some very ancient pseudogenes or retroposon insertions may have undergone so many random mutations since their insertion that their identities as pseudogenes or retroposons have been obliterated. However, at the rate of mutation estimated for nonselected sequences, complete obliteration of a typical retroposon by mutations would require over 100 million years. Hence it would not be surprising to an evolutionist that functionless retroposon sequences that inserted into a common ancestor of humans and cows might still be detectable by computerized comparisons of DNA sequences.
5.4 Important roles have been found for DNA regions previously thought to be functionless
At a recent debate with me Dr. Gish cited a review in Science entitled "Mining treasures from 'junk' DNA" (263:608, 1994), seeming to imply that this review suggests functions for pseudogenes and retroposons that would be consistent with the creationist view that they were designed to function similarly in similar species. In fact, this review discusses evidence for possible functions of centromeric and telomeric repetitive sequences, minisatellites, introns and 3' untranslated regions. It mentions pseudogenes and retroposons but makes no suggestion that these particular elements have function, so this review offers no argument against the points made in this essay. Nevertheless, since there have been other speculations about possible functions for DNA outside gene coding sequences, it is worth considering why scientists generally accept the notion that most of this DNA is junk.
First, we know several mechanisms by which DNA length can be increased through genetic accidents such as DNA duplications and insertion of retroposons, which have been observed in the lab or occurring in humans without apparent effects; so it is reasonable to suppose that these mechanisms operated in the past to increase genome size without affecting function. There appears to be little or no selective pressure to reduce the size of vertebrate nuclear genomes; and there is no apparent mechanism to selectively eliminate useless DNA. Large deletions that eliminate functional DNA are selected against. These observations would predict the accumulation of useless DNA as the result of random genetic accidents, so when we see DNA that seems non-functional, we shouldn't necessarily assume that it has function that we don't understand.
Second, when DNA sequence is compared between species like human versus mouse, sequences that are known to have function -- coding sequences of genes in particular -- are found to be highly similar, consistent with selective pressure that weeds out individuals that have deleterious mutations in these functional regions. Conversely, DNA regions with no known function -- e.g. non-coding sequences between genes -- generally behave as if they are under no selective pressure, that is they apparently accumulate mutations at a much higher rate so there is little sequence conservation between distantly related species. As an exception that probes the rule, comparisons of non-coding sequence across species occasionally detect "islands" of short conserved sequence in non-coding regions. Some of these have turned out to correspond to regulatory regions like promoter or enhancer elements that control when a nearby gene is expressed. An example of such an "island" conserved between rabbit, mouse and human was discovered in my own lab [Emorine et al., Nature 304:447, 1983]; it turned out to represent an important enhancer. These kinds of regulatory regions generally take up much less DNA than the coding sequences of the genes they regulate, so they cannot represent a likely function for most non-coding DNA. The good correlation between function and sequence conservation lends support to the idea that most poorly conserved sequences do not have function. However, it should be noted that for most of the "islands" of conserved sequence in DNA between genes (Shabalina et al., Trends Genet 17:373, 2001), no function has yet been discovered. Some may include RNA species that function without being translated into protein.
A third but related argument derives from the observation that the insertion of a retroposon into a functional sequence is a potent way to destroy that function. Examples of naturally occurring insertions were discussed in section 5.2 above; and intentional retroposon insertion is being widely used as a laboratory tool to create panels of mouse, drosophila or yeast strains with different gene functions destroyed. However, most examples of retroposon insertions between genes do not have any apparent affect on individuals harboring them; for example the Alu sequences that are polymorphic in human DNA appear to be harmless when present. Therefore, it is reasonable to infer that these insertions did not interrupt any functional sequence. (Of course it is impossible to rule out the formal possibility that some hypothetical functional sequences outside genes can still function despite the presence of a retroposon insertion.)
Finally, several examples are known of pairs of species that have similar apparent complexity but widely different genome size (C-value paradox). The pufferfish Fugu has about one fourth the genome size of other fish species but about the same number of genes. The main difference is a smaller amount of DNA between genes in Fugu DNA (e.g. see Elgar et al. Genome Res 9:960, 1999). Although questions remain about the interpretation of this difference, it would seem that much of the DNA between genes in most fish genomes (and probably in ours also) is dispensable. (Conversely, the small regions of non-coding sequence that are conserved between Fugu and Homo frequently correspond to functional regulatory sequences.)
It is impossible to prove absence of function for any region of DNA. Moreover, it is likely that some function may be found for a few additional short regions of non-coding DNA that are not currently recognized to have function. Nevertheless, as indicated above, scientists draw tentative conclusions based on data currently at hand rather than on hypothetical possibilities of future data; and the arguments I just presented based on presently available evidence suggest that most DNA sequences that appear to be functionless are just that.
5.5 Pseudogenes serve a function: they provide a "backup" copy that can be corrected to encode a useful protein if the functional gene gets critically mutated.
Dr. Gish provided no specifics for this claim, but perhaps he was referring to a recent suggestion that a bovine seminal ribonuclease pseudogene was recently "corrected" to become functional by a process known as "gene conversion" (Trabesinger-Ruef et al. FEBS Lett 382:319, 1996). Although this may occasionally happen, far more instances have been described in the literature in which defects in a pseudogene induce damaging mutations in a nearby functional gene by gene conversion, inactivating the functional gene. As an example of this kind of event, in almost all patients suffering from deficiency in steroid 21-hydroxylase because their (normally) functional 21-hydroxylase "B" gene copy has been inactivated by point mutations, these mutations apparently resulted from gene conversion by the "A" pseudogene copy (Collier et al, Nat Genet 3:260, 1993; Carrera et al. Hum Hered 43:190, 1996). Similar gene conversions by a pseudogene are thought to have inactivated the glucocerebrosidase gene in Gaucher disease patients (Eyal et al. Gene 96:277, 1990), the gene 14.1 encoding an immunoglobulin "surrogate light chain" in a patient with immunodeficiency (Minegishi et al., J Exp Med 187:77, 1998), the ABCC6 gene in patients with pseudoxanthoma elasticum (Cai et al, (J Mol Med 79:536,2001), the PKD1 gene in autosomal dominant polycystic kdney disease (Watnick et al. Hum Mol Genet 7:1239, 1998), the beta-crystallin B2 gene in autosomal dominant hereditary cataract (Vanita et al. J Med Genet 38:392,2001), the NCF1 gene in chronic granulomatous disease (Vazquez et al., Exp Hematol 29:234,2001) and the von Willebrand factor gene in patients with von Willebrand disease (Eikenboom et al., PNAS 91:2221, 1994), to name just a few examples. In other cases gene conversion events have apparently transferred genetic information between two pseudogenes (Shapiro and Moshirfar, J Mol Biol 209:181, 1989) or between two functional genes (Ollo and Rougeon, Cell 32:515, 1983). Because gene conversion involving pseudogenes has been reported to occur with harmful or neutral effects more than it has with beneficial effects, the hypothesis that pseudogenes were "designed" with the potential for gene conversion as their purpose seems unconvincing. (The one example where pseudogene copies clearly do fulfill an important function in transferring their sequence to another gene copy by gene conversion occurs in the somatic diversification of immunoglobulin variable region genes of chickens and rabbits; of the many mutations that are generated by this mechanism, those few that provide a "better fit" between the immunoglobulin and its target antigen are selected for expression. This selection for improved function among genes that have undergone quasi-random sequence changes is an attractive biological model for the evolutionary improvements in protein function. Ironically, in several debates with me Dr. Gish denied that such somatic diversification occurs, although he was obviously totally ignorant about the scientific literature concerning antibody genes.) In addition to being unconvincing for the reason described above, Dr. Gish's idea that pseudogenes were created to provide a "backup" gene copy offers no creationist explanation for the more numerous shared retroposons that are not pseudogenes.
5.6 All this retroposon stuff is really too hard to understand.
Dr. Gish used this appeal to the audience at a recent debate with me. He seemed to be coaxing the audience to ignore the implications of the argument from shared pseudogenes and to disregard the fact that he (Dr. Gish) could not find valid counter-arguments to oppose it. This is a typical debate maneuver for creationists: using humor or invocation of faith or some other irrelevant appeal to distract a lay audience from realizing that a creationist position has been effectively refuted.
(This essay was sent to Dr. Gish to solicit any further arguments against the points made here. No reply was received.)
5.7 In addition to Dr. Gish, creationist John Woodmorappe has commented on pseudogenes (Noah's Ark, a Feasibility Study, 1996, published by ICR, p. 202; Bible-Science News 33:7,1995). He makes several of the same arguments as Dr. Gish (see 5.2 and 5.4 above) but adds a few of his own. A creationist interpretation of pseudogenes offered by Woodmorappe is that some pseudogenes may be "the result of degenerative changes in living organisms since the Fall." This interpretation seems plausible, and--if we ignore the "Fall" part--not very different from the evolutionary idea that pseudogenes arise by random genetic accidents. However, this interpretation completely ignores the fact that many pseudogenes are shared between apes and humans, located in the same positions and sharing the same genetic defects, apparently the result of the same genetic accident or "degenerative change" in a common ancestor. (If these shared pseudogenes arose after the "Fall" as suggested by Woodmorappe, did the "Fall" perhaps occur before man diverged from the apes?)
5.8 In addressing shared pseudogenes, Woodmorappe tries to cloud their strong support for evolution by claiming that for particular pseudogenes the degree of "relatedness" inferred from the presence or absence of the pseudogene in different species contradicts the species "relatedness" inferred by evolutionists from other characteristics. In this argument, Woodmorappe falls in line with other Creationist arguments that invite us to discard evolution because of specific cases that violate a simplistic interpretation of evolution, and to ignore the vastly greater number of examples that support evolution. In the long and complex history of life on earth, many exceptions to simplistic notions have been generated--e.g. cases where older fossils lie above younger ones (because of folding of geologic strata or thrust faults) or examples where sequence similarities of small stretches of DNA compared between species seem to violate accepted relationships (because of statistically expected errors due to small samples). Similarly, we can expect cases in which a pseudogene or retroposon that arose in the ancestor of three modern species (A, B and C) may get deleted in one (say C), suggesting a closer relationship between A and B than is warranted on other grounds. An example like this should not cause us to discard what we learn from the majority of shared pseudogenes and retroposons; rather, we should use caution in drawing generalizations from exceptional cases.
| Box 3 |
|---|
|
Woodmorappe describes an example of an epsilon immunoglobulin pseudogene that was reported (Ueda et al, PNAS 82: 3712 1985) to be shared by gorilla and man but not by chimpanzee, seeming to contradict the conventional evolutionary view that human ancestors diverged from the gorilla lineage before they diverged from the chimpanzee lineage. Unfortunately, Woodmorappe failed to consider later data from Ueda's laboratory (Kawamura and Ueda, Genomics 13:194, 1992) that were available when Woodmorappe wrote in 1994 (Bible Science News 32:4 p. 12). These more recent data show that DNA deletions destroying duplicated copies of the epsilon immunoglobulin genes (1) occurred independently in human and gorilla lineages (independence was deduced from the fact that the "right" and "left" boundaries of the deleted DNA were completely different in the two species), and (2) also occurred (again independently) in chimpanzee. Thus Woodmorappe's example of a shared pseudogene linking humans to gorilla but not to chimp (in apparent violation of the more recent divergence of human ancestors from chimpanzee accepted by most evolutionists) is incorrect: these are not "shared" pseudogenes but independently arising pseudogenes, and chimpanzee has a similar, though larger, deletion. (I should mention that I cited this same incorrect example in my original version of this essay. However, at the time I wrote--1986--the example was supported by the evidence then available; and I printed a correction in Creation/Evolution after the new data were published. I should also stress that the example of the processed epsilon pseudogene mentioned in section 4.3 above represents a completely different sequence, which no one disputes is shared by humans, chimps and gorillas.) |
However, the example of shared pseudogenes that Woodmorappe offers to challenge the evolutionary model has more mundane explanation: it is simply based on outdated incorrect information (see box 3).
5.9 A final hypothesis offered by Mr. Woodmorappe
(in personal correspondence) is that similar genomes (like
those of human and chimp) might tend to acquire the same
pseudogenes independently, while less similar genomes may
be less able to acquire the same pseudogenes. This
obviously ad hoc hypothesis would theoretically explain
why--even if humans and chimps were independently
created--they might share more pseudogenes than less
similar, independently related species pairs such as human
and gibbon. The problem with this hypothesis is that
independent occurrence--i.e. in two different
individuals--of the same retroposon inserting at the same
position has almost never been reported, even in
individuals of the same species. I have been able to find
only four publications describing examples of identical
independent insertions. One involves a modified Rous
sarcoma virus engineered with a specific selectable marker
and infecting turkey fibroblasts grown in tissue culture
(Shih et al, Cell 53:531, 1988); and even in this unusual
paper with a specially engineered virus the frequency of
such insertions was estimated at only 1 in 4000 insertion
events. The second example is a very recent and
controversial publication (Slattery et al., Mol Biol Evol
17:825, 2000) which interprets two identical insertions of
a SINE at the same location (an intron of the gene Smcy) in
a domestic cat and a bobcat as representing independent
insertions rather than reflecting a single insertion in a
common ancestral feline. Two additional publications (Kass
et al, J Mol Evol 51:256 2000; Cantrell et al., Genetics
158:769, 2001) describe apparent identical but independent
insertions of SINEs in mouse species. (John Woodmorappe
declined to cite any data at all when challenged to provide
examples of independent insertions. However, if readers of
this essay are aware of other evidence for independent
insertions of the identical element at the identical
position in any laboratory models, I would appreciate
appropriate citations and will update this essay to reflect
them.) Very many naturally occurring insertions have been
documented in yeast TY elements, drosophila gypsy and P
elements, murine retroviruses and transgenes, and human HIV
insertions--all without identical independent insertions
having been reported. If independent organisms of the same
species (i.e. with genomes more nearly identical than human
versus chimp) almost never acquire the same pseudogene or
retroposon insertion at the same position, it is hard to
take seriously the hypothesis that, for example, the same
seven Alu inserts in same positions of the human and
chimpanzee ![]() globin locus (see
section 4.4 above) could have occurred as 14 independent
insertion events.
globin locus (see
section 4.4 above) could have occurred as 14 independent
insertion events.
5.10 Couldn't a pseudogene have been transmitted by a virus from one species to another, leading to shared pseudogenes? A proposal along these lines has been suggested by anti-evolutionist Pat Kohli and seems superficially plausible. Several viruses, including retroviruses, are known to occasionally pick up nucleotide sequences from a "donor" cell which can then, after reinfection of a new cell, be inserted into the DNA of the new "recipient" cell. Indeed this mechanism is known to have significant consequences: if the transmitted DNA includes a mutated version of certain key genes regulating cell division, such a DNA sequence can act as an oncogene and cause malignancy in the recipient cell (Bishop, Cell 42:23, 1985). Theoretically, a pseudogene or retroposon sequence might captured by a virus and then be transmitted across species by this mechanism, leading to the existence of identical useless sequences shared between two species. Indeed, rare instances of apparent cross-species transfer of retroposons have been reported (e.g. between two fruit fly species [Jordan et al, PNAS 96: 12621, 1999] or from venomous snake to ruminants [Kordis and Gubensek, PNAS 95:10704, 1998; Kordis, Genetica 107:121, 1999]). However, this is not likely to be the explanation for most shared pseudogenes/retroposons for at least three reasons.
First, shared pseudogenes/retroposons are generally found at the exactly homologous position in the DNA from each species. This is almost always true in the case of classical pseudogenes, which lie in close proximity to the functional gene, and it is also true for Alu sequences like those mentioned in the globin gene cluster, and for processed pseudogenes whose location has been determined (e.g., the human immunoglobulin epsilon processed pseudogene mentioned above [Ueda, et al, EMBO J 1:1539, 1982; Tanabe et al. Cytogenet Cell Genet 73:92, 1996]). Target sites for viral insertion may share certain local sequence features (Craigie in Trends in Genetics 8:187, 1992; Knoblauch et al., J Virol 70:3788, 1996; Stevens and Griffith, PNAS 91:5557, 1994), but these features occur quite frequently and are generally scattered throughout the recipient cell DNA. Other than the papers mentioned in section 5.9, there is no precedent or known mechanism for a virally transmitted DNA segment to target a specific location in recipient cell DNA, as would be necessary for a pseudogene representing a hypothetical viral insertion to occur at the same location as the hypothetical donor sequence. Therefore, the shared locations of pseudogenes/retroposons with respect to surrounding DNA argue strongly against such a model of cross-species transmission.
Secondly, if most shared pseudogenes/retroposons represented virally-mediated transfer from one species into another, one would expect to find viral sequences near pseudogenes in "recipient" species. Such viral sequences are regularly present in known examples of viral transmission of DNA from one cell to another, including insertions of engineered retroviral constructs; but viral sequences are not found associated with most pseudogenes/retroposons other than endogenous retroviruses.
Finally, several genealogic trees have been generated by comparing across species for the presence or absence of LINE or Alu insertions at specific locations in the genome, an exercise similar to that shown in Figure 5 above (Malik et al, Mol Biol Evol 16: 793, 1999; Hamdi et al J Mol Biol 284:861, 1999); the retroposon-derived genealogic trees were precisely congruent to trees previously established based on sequence similarities and anatomic features. If cross-species tranfer explained most shared retroposons, no such congruence would be expected.
In a computer-assisted search of the scientific literature, I could find only two examples of pseudogenes for which viral transmission was even tentatively considered as a mechanism of origin, in both cases with rather weak evidence (Gruskin et al., PNAS 84:1605, 1987 and Robins et al., J Biol Chem 261:18, 1986). Readers who are aware of other examples are invited to Email them to me for inclusion in future updates of this article. For the present, the evidence argues against virally-mediated cross-species transfer as a general mechanism to explain shared pseudogenes/retroposons.
5.11 Creationist L. J. Gibson has also addressed pseudogenes in a published article (Origins 21:91, 1994). Gibson's article boils down to two points, one similar to that discussed in section 5.2 above, plus an additional more philosophical point. He notes that the argument from shared pseudogenes rests on the assumption "that God would not create similar non-functioning sequences in separate species," which he calls "a theological argument [which] can hardly be addressed by science" and which would require Scriptural support to be believed. Since the Bible does not address--and therefore leaves open--the possibility that God might create non-functional sequences in DNA, Gibson feels that one cannot dismiss the notion that God did in fact create such sequences individually as he created each species, including those non-functional sequences that we now find shared between different species.
It should be mentioned parenthetically that Gibson's argument undercuts the creationists' own interpretation of species similarity mentioned at the beginning of this essay (section 1.2). As we discussed, creationists have claimed that the similarity trees based on sequence information need not be accepted as evidence for evolutionary relationships, because species independently created by an intelligent designer might be expected to show identical patterns of apparent relatedness. Gibson's criticism applies equally well against this creationist argument, as the Bible does not mention God's plans for sequence similarity.
However, as we discussed earlier, this creationist notion of similar sequences designed for similar functions at least makes some intuitive sense. In contrast, Gibson proposes a clearly unacceptable ad hoc hypothesis when he suggests that a designer might have placed non-functional retroposon insertions--mimicking all the features of those currently retroposing randomly in the laboratory--into the same positions of independently created species' DNA; this idea merits as much credence as the claim that shared false entries in a directory are due to independent mistakes rather than plagiarism. Gibson's hypothesis does not argue for a creator making understandable design decisions, but a creator so unpredictable that he could be the author of any scientific findings traditionally interpreted as undesigned--unless the Bible specifically states otherwise. Thus Gibson's logic would support the following statements, because they are not specifically contradicted by the Bible: (1) God created fossils looking like the remains of animals who never lived, and embedded them in rocks. (2) God created radioactive elements in rocks that would falsely suggest ages older than their actual ages. (3) God created the universe 6000 years ago with starlight on its way to our eyes but with the properties expected of light that left stars billions of years ago. In other words, Gibson's logic invites us to reject any scientific argument for evolution if that argument is not specifically verified in the Bible. Gibson's view may be internally consistent, but it clearly requires that the truth of the Bible be accepted on faith as a basis for judging the merits of scientific conclusions, and thus it departs from true science based on hypothesis testing, inductive logic, and conclusions based on observed data. If a scientist sees a retroposon inserting in the laboratory as a result of several known biochemical parameters, Occam's razor discourages him from postulating an intelligent hand guiding its creation. If we find other insertions in our DNA with identical features to those arising under observation, we assume that the ones in our DNA arose by a similar mechanism. We know that such insertions arising under laboratory observation can be used to trace the lineage of laboratory animals, and that other natural insertions can be used to trace populations in the wild; we have no reason to dissuade us from using similar insertions to trace the lineages of different species. This inductive reasoning is fundamental to paleontology, radioactive dating, astronomy, physics, medicine and every other field of science. If Gibson feels that the merits of a scientific argument depend on how well it is supported by the Bible, he can simply dismiss evolution outright because it conflicts with Genesis, and avoid the bother of dealing with all the details of the individual scientific observations and deductions on which evolution is based. This "Bible first" approach may be appropriate for religion, but it is unacceptable as science.
6. Testing the model
One feature of science that distinguishes it from revealed religious belief (and evolutionists from creationists) is the scientific conviction that new knowledge about the past can be obtained from thoughtfully designed analysis of the modern world. Creationists often claim that, since the origin of species occurred in the distant past, there is no scientifically valid way to study the process today and so evolution is not real science testable by experiment. However, even without actual experiments, a scientific hypothesis can be tested if it suggests a non-trivial prediction that can be verified, or falsified, by the collection of more data.
Indeed the interpretation of shared processed pseudogenes outlined here represents a hypothesis that can be tested because it presents a rather startling implication: from a comparison between two nucleotide sequences from a single species--that is, the sequences of a processed pseudogene and the functional gene from which it derived--it should be possible to predict which other species will share the same pseudogene and which will not. To understand the logic of such a prediction, consider the fact that if a processed pseudogene arose in an ancient species, copies of that pseudogene should be found in the modern descendants of that species. Thus, according to the evolutionary model, if we knew when a human processed pseudogene arose, and could thus fix its origin to a particular position on the accepted evolutionary "tree," we would predict that the same processed pseudogene should be found in modern species that derive from that point on the tree and not in any other branches.
In fact, there is a way to estimate when a given processed pseudogene was formed. It turns out that "silent" mutations--that is, mutations that have no effect on the survival of the organism (like all mutations in useless pseudogenes)--accumulate at a fairly uniform rate. This rate has been estimated by examining the number of "silent" sequence differences between corresponding functionless sequences in two species and comparing this number with the approximate date of divergence of the same two species as indicated by the fossil record. Given this mutation rate and the number of sequence differences between a particular processed pseudogene and its functional source gene (from the same species), one can estimate the date of origin of the pseudogene; then, based on this date, one can derive predictions about which other modern species should carry the same pseudogene. These predictions can be tested by searching for the pseudogene in a variety of species.
Consider, for example, the processed human epsilon pseudogene discussed earlier (section 4.3). The number of nucleotide differences between this pseudogene and the functional gene suggests that this pseudogene arose about 40 million years ago. Therefore, the evolutionary interpretation of processed pseudogenes presented in this essay would predict that mice and rabbits (which are thought to have diverged from the human lineage 70 to 80 million years ago, before the apparent origin of the pseudogene) should not carry this pseudogene. In contrast, apes and Old World monkeys--whose estimated dates of divergence from the human lineage (5-10 and 30 million years ago, respectively) are both after the apparent pseudogene origin--would be expected to carry the pseudogene. Available evidence confirms all of these predictions and is also consistent with similar predictions about the species distribution of other processed pseudogenes (see for example Anagnou et al. PNAS 81:5170, 1984 with respect to dihydrofolate reductase, Craig et al. Gene 99:217,1991 with respect to triosephosphate isomerase, and Friedberg and Rhoads Molec. Phylogenet & Evolution 16:127, 2000 with respect to enolase, calmodulin and argininosuccinate synthetase).
By a similar logic, it is possible to estimate the age of insertion of an endogenous retrovirus by comparing the sequences of the "left" and "right" LTR (see section 2.2.2.d above). Since the "left" LTR is copied from the "right" LTR sequence at the time of insertion, the two LTRs share identical sequence at the time the retrovirus copy originates. After insertion, the two LTRs accumulate mutations independently, and so the number of sequence differences between the two LTRs can be used to estimate the age of a particular retroviral insertion; this age can be used to predict the species distribution of shared copies of the particular retrovirus insertion. When the ages of several human endogenous retroviruses were estimated recently using this approach, the predicted species distribution of shared copies was confirmed (Johnson & Coffin, PNAS 96:10254, 1999).
While individual human SINE and LINE insertions cannot easily be dated by sequence analysis alone, it may be possible to estimate a rough time period when certain subclasses inserted. This is possible because specific classes of similar retroposons are thought to have populated the mammalian genome in waves, with certain families (and subfamilies, in the case of Alu and LINE sequences) being copied from a small number of source retroposons active at any particular period in our evolutionary history. This model has been deduced from the fact that for some families of human retroposons (e.g. LINE2), comparisons between specific members of the family reveal relatively divergent sequences, as though individual copies have accumulated many different mutations over a long time since inserting into our DNA (Nature 409:860, 2001, see p 881), while others, such as the Alu sequences (particularly the Ya5 and Ya8 subfamilies) and the Ta family of LINE1 sequences, show fewer deviations from a consensus sequence and are therefore thought to have inserted more recently. These evolution-based interpretations of human retroposon sequences predict wide species sharing of putatively older retroposons but more restricted sharing of putatively younger ones, and these predictions are confirmed by independent species distribution data for these retroposons (Gonzalez et al, Genomics 18:29, 1993; Shaikh and Deininger, J Mol Evol 42:15, 1996; Carroll et al, J Mol Bio 311:17, 2001; Sheen et. al. Genome Res 10:1496, 2000; Boissinot et al., Mol Biol Evol 17:915, 2000).
More shared retroposons will certainly be discovered, and only time will tell how consistently evolutionary predictions like these are confirmed. But at present, almost all available data are consistent with evolutionary models of common descent, and no alternative creationist rationale for explaining this consistency has been proposed. Repeated instances of this kind of prediction and confirmation can supply convincing evidence for evolution even if some kinds of direct experiments, like studies on living dinosaurs, are impossible. (If readers are aware of other examples of processed pseudogenes or other retroposons whose distribution in different species either supports or contradicts the accepted "genealogic tree" I would appreciate hearing about these cases by Email, and would revise this posting as appropriate.)
7. Conclusion
Do the shared functionless sequences described here prove that humans and apes had a common ancestor? Actually, no scientific knowledge is based on unassailable proof of the sort that supports mathematical theorems, so the creationist complaint that evolution has "never been proven" simply reveals a gross misunderstanding of the nature of science. Rather, science advances by the accumulation of clues sought by persistent detectives (scientists) who try to derive logical and unbiased deductions from these clues. Like a jury presented with these clues, we can try to arrive at the most likely verdict even though we recognize that our facts are incomplete; there are no living "witnesses" to the eons of evolution, so we must make the best deductions we can from the clues at hand. In "the case of the shared functionless sequences," an unbiased jury would surely conclude that copying from a shared ancestor was the most likely explanation, consistent with the evolutionary interpretation. This conclusion would follow the logic of actual copyright law in which shared errors are accepted as evidence of copying. The strong acceptance of this conclusion among scientists is indicated by the fact that no alternative explanation has been proposed in the scientific literature to explain the widespread sharing of so many functionless sequences between species. Thus, if we are to accept the evidence of science, it would appear that common descent of disparate species from a shared ancestor ("macroevolution" in the creationist terminology) has actually occurred.
As new examples of shared pseudogenes and retroposons are discovered by molecular geneticists, this information will join the immense body of clues from other disciplines which, collectively, already provide overwhelming evidence for evolution. Despite this impressive evidence, no scientist believes that all the answers are in on evolution or that our current understanding of pseudogenes and retroposons is immune from revision in light of future knowledge. Indeed, scientists in laboratories throughout the world are continuing to probe the genes of various species, comparing the molecular genetics data with the fossil record and refining our knowledge of the history of our species.
At the present stage of this never-ending research, the
evidence suggests what to me is an awesome notion: like a
biological Rosetta Stone or Dead Sea Scroll, our own
DNA--an Encyclopedia Brittanica's worth of information in
every cell of our body--contains a record of the past which
we are just now learning to read. This record, reflecting
millions of years of genetic history, includes the relics
of ancient genetic accidents that occurred before our
ape-like ancestors roamed the plains of Africa, relics that
we now share with other descendants of those same
ancestors: modern gorillas and chimpanzees.
| The FAQ | Must-Read Files | Index | Creationism | Evolution | Age of the Earth | Flood Geology | Catastrophism | Debates |